On January 9, 2007, Steve Jobs launched a revolution when he pulled an unfamiliar device out of his pocket and told a room of 5,000 tech-geeks that his company was “going to reinvent the phone.” The device could do unimaginable things, he said, like search the web, make calls, and play music all with a remarkably simple user-interface. At the time it did not seem possible, now it’s hard to imagine life without the iPhone and its descendants.

Most end users of technology tend to accept the status quo until they’re handed something better—its human nature. But researchers at the Novartis Institutes for BioMedical Research (NIBR) are actively pursuing disruptive technology before it takes them by surprise. Teams are scouring academic labs for new inventions and launching collaborations to push the development of equipment in a direction that will benefit scientists working in drug discovery.
A study published online in Nature Chemistry on May 25 is the culmination of one such collaboration. Researchers used a novel imaging technique invented at Harvard to visualize the small-molecule drugs imatinib and nilotinib inside the cell without using molecular tags.
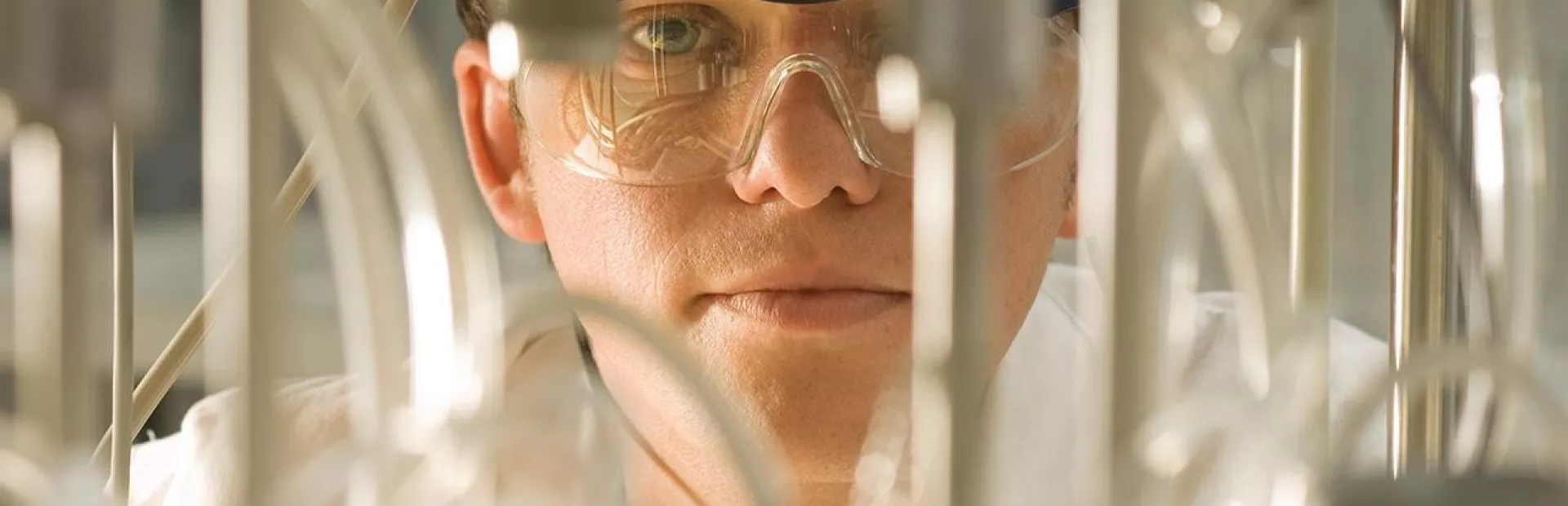
In drug discovery, scientists might get annoyed with the limitations of a particular microscope or mass spectrometer, but they typically suppress such feelings, bottling up frustration for the sake of the experiment in progress. Steve Martin, Head of Analytical Sciences and Imaging at NIBR, is pushing his teams to harness their malaise to improve—and even transform—lab equipment.
“If we’re going to cure patients five years and ten years from now, then we’re going to have to evolve the technology we use to understand biology and the chemistry,” he explains. “We can’t just take what’s on the shelf and use it as it is. That’s just not going to get us where we need to be.”

To “get disruptive” Martin and other leaders have convened groups (see sidebar) of biologists and chemists to discuss their technology needs, urging them to dream big. He suggests that they suspend disbelief during these conversations. In a lab without limits, what types of molecular properties or behaviors would they measure or visualize? Some requests can be addressed through incremental improvements to tools on the market; others require looking beyond existing products to technology on the horizon.
One such conversation led NIBR scientists to realize that they needed better ways to analyze the behavior of drugs inside cells.
“It’s a central question for scientists in drug discovery to see how our molecule is able to function inside the cell,” says co-first author Jing Zhou, Senior Investigator I in Analytical Sciences and Imaging. “It’s also one of our dreams to see it in real time in a living cell in as natural a state as possible.”
But when it comes to small molecules—whose structure and function can fundamentally change when tagged with an imaging label—that dream was not yet a reality. So Zhou turned to the scientific literature for ideas.
He gave himself some constraints: find a novel technology that would allow him to see small molecule drugs inside a living cell, in real time, without labels, and at subcellular resolution. After sifting through hundreds of academic papers, a study by Sunney Xie, Professor of Chemistry and Chemical Biology at Harvard University, on a technique called raman spectroscopy was the last one standing.
Raman spectroscopy uses a laser beam to detect the vibrational signature of a molecule, which is generated by its chemical bonds. Imagine a satellite tracking a GPS device around a city. The GPS device, like the molecule in question, sends out a unique signal, which the satellite is able to distinguish from everyone else on the road to determine exactly where you are on the map.
But traditional raman spectroscopy isn’t very sensitive. Xie’s team solved the sensitivity problem by adding a second laser beam at a second frequency to significantly increase the acquisition speed. Zhou wanted to test the refined technology with some of the company’s small molecule drugs, so he reached out to Xie, who quickly agreed to collaborate.
“We’ve been developing this for more than a decade, and I’m very interested in using it to solve real-world problems,” says Xie, who is the senior author on the Nature Chemistry paper. The team decided to focus on two well-known oncology drugs. “It was kind of surprising to me that there wasn’t a very good way of looking at the distribution of these drugs inside the cell.”
The team showed that many imatinib and nilotinib molecules get trapped in small compartments within cells called lysosomes, which function like garbage disposals, breaking down “waste” from the cell. Cancer cells seem to defend themselves from therapeutic attack by quarantining these drugs, keeping them away from their intended targets. But the researchers were able to reduce the concentration of imatinib in lysosomes by co-administering it with a molecule called chloroquine. Apparently, the chloroquine allows imatinib to leak out of the lysosomes.
Partnerships like this, Martin says, will enable drug discovery scientists to guide the development of instrumentation that will most benefit their work. Zhou agrees. “We can help frame and repurpose the technology that’s been developed in all kinds of academic labs,” he says.
The following NIBR scientists are authors on the Nature Chemistry paper: Jing Zhou, Wenjing Suzanne Zhu, Paul W. Manley, Y. Karen Wang, Tami Hood and Andrew Wylie.